Liver projects
High resolution imaging of liver tissue
People: Hidenori Nonaka, Jerome Gilleron, Anja Zeigerer, Pablo Amaya
The goal of our analysis is to understand the complex organization of the liver tissue. Hepatocytes, the most abundant cell type in the liver, display a peculiar polarity characterized by multiple apical surfaces, organized into bile canaliculi, having a complex three-dimensional organization to collect and transport bile. Endothelial cells also organize themselves accordingly by arranging an intricate network of sinusoids around hepatocytes to allow efficient nutrient exchange with the blood. This organization is fundamental for the liver to implements two of its main functions: bile secretion and blood filtering.
To be able to understand the design principles of liver organization and accurately describe its morphology we cut and stain by immunofluorescence thick liver samples, we visualize them by confocal imaging and we analyze them by automated image analysis. Our strategy is to understand liver morphology and function by accurate and quantitative measurements of different structures in the liver: bile canaliculi, sinusoids and hepatocytes. We are also interested in zonation properties within liver lobules: bile and blood have directional (and opposite) flows between central and portal veins. The understanding of how this is achieved in terms of tissue organization is very interesting and of relevance for liver physiology.
Additionally, we are also interested in understanding how tissue is built during development through the analysis of fetal liver: images for the quantitative comparison of fetal and adult liver are also acquired. Interesting in vitro systems, such as primary isolated mouse hepatocytes, isolated hepatoblasts (precursor cells) and co-cultures of hepatocytes and endothelial cells, will also help to understand principles of hepatic cell polarity and tissue organization: imaging-based approaches and quantitative analysis of cellular and sub-cellular structures have being applied also for these systems.
In the future, we will extend our analysis to sub-cellular structures, in particular endosomes, to understand the organization of the endocytic system in liver cells and its contribution to cell polarity and tissue morphogenesis.
Image analysis of liver tissue
People: Giovanni Marisco, Hidenori Nonaka, Yannis Kalaidzidis
To realize our goal of understanding liver architecture and function we perform image analysis of thick liver samples. To do this, we develop customized methods for image processing and feature extraction for the quantitative description of different morphological features. The algorithms implemented are integrated into a software suite developed in our lab, called MotionTracking: the use of in-house built software has the advantage of providing the necessary flexibility for new solutions of the challenging tasks that biological research poses every day.
Several procedures have been developed for this purpose and are visually summarized in the figure: here, the analysis of bile canaliculi network is shown, but the processing of the endothelial cell network and hepatocyte cells is conceptually similar. Background subtraction and thresholding aims at identifying objects of interest in the image: the task is particularly challenging for bile canaliculi, given their complex connectivity and the relatively small size (also below 1 µm in radius). The algorithm developed is robust and sensitive enough to detect most of the structures of interest, to preserve the original connectivity of the network and to efficiently remove background area (upper left part in the mosaic image). Surface rendering with triangulation methods allows to smoothly represent the contour of the segmented objects on the binarized images, and to estimate the surface area and the inner volume of tubes (upper right part in the mosaic image). The connectivity of bile canaliculi into a network is of high relevance for the analysis of morphology in the liver: for this, a procedure that estimates central lines and represent them as connected graphs has been developed (middle and lower right part in the mosaic image). The procedure can be applied as well to the analysis of the sinusoidal network. Compared to classical approaches such as skeletonization, this procedure allows a better pruning of spurious branches deriving from small structures (e.g., staining artifacts, processing errors or small uncertain protrusions).
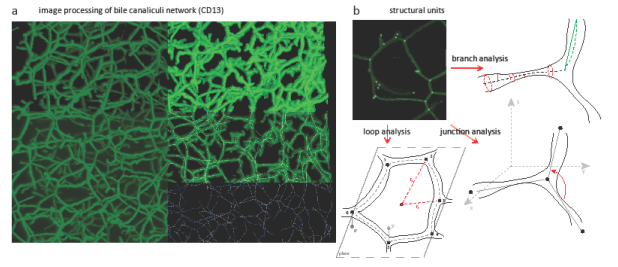
Image processing and analysis of liver tissue. a) Montage image obtained through the projection of 3D stacks displaying several steps of image visualization and processing: background subtraction, thresholding, surface rendering, and central line extraction. b) Image analysis for the identification of structural units in the bile network: descriptors for quantifying the geometrical, orientation and spatial features of branches, junctions and loops are designed and implemented.
Image processing is a fundamental and challenging task in the analysis of tissues, but to identify the minimal structural units and identify the design principle of liver tissue organization is necessary to extract quantitative parameters. Feature extraction is this process of obtaining information from images. A panel of statistical descriptors has been designed and developed to this purpose: different geometrical, orientation and spatial properties of the network are calculated. Three main categories are analyzed by those: branches, junctions and loops, which taken together constitute the structural elements of tubular network (both canaliculi and sinusoids).
On the technical side, many of the computationally intense procedures are translated to run on the GPU instead of the CPU, in order to take advantage of the massive parallelization that graphic card computing allows. Image processing is naturally suited for high parallelization, since many algorithms perform the same routine on each one of the millions of pixels (or voxels) in the image.
Future work is required for the optimization of methods for the segmentation of the endothelial cell network and the identification of hepatocyte cell borders, and for the tiling of several image stacks for covering big tissue areas. New algorithms and statistical descriptors are also required to correlate the information belonging to cell, sinusoids and bile canaliculi. Finally, the development of new methodologies for the integration of information at different scales (tissue level, cell level and sub-cellular levels) represents a challenging task and an active field of our future research effort.
Collaborators: Jan Hengstler (Dorthmund) for the experimental procedures; Carsten Werner (Leibinz) for bio-engineerd substrates; Lutz Brush and Walter de Back (ZIh) for the modeling.